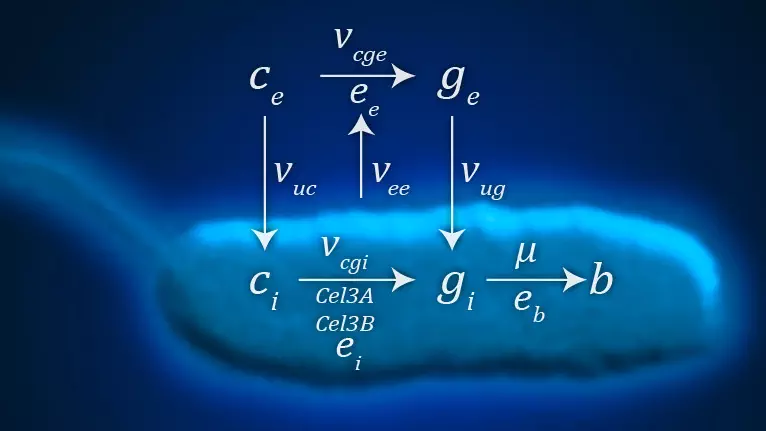
Systems Biology
We build quantitative mathematical models to understand, analyze, and predict the behavior of biological systems.
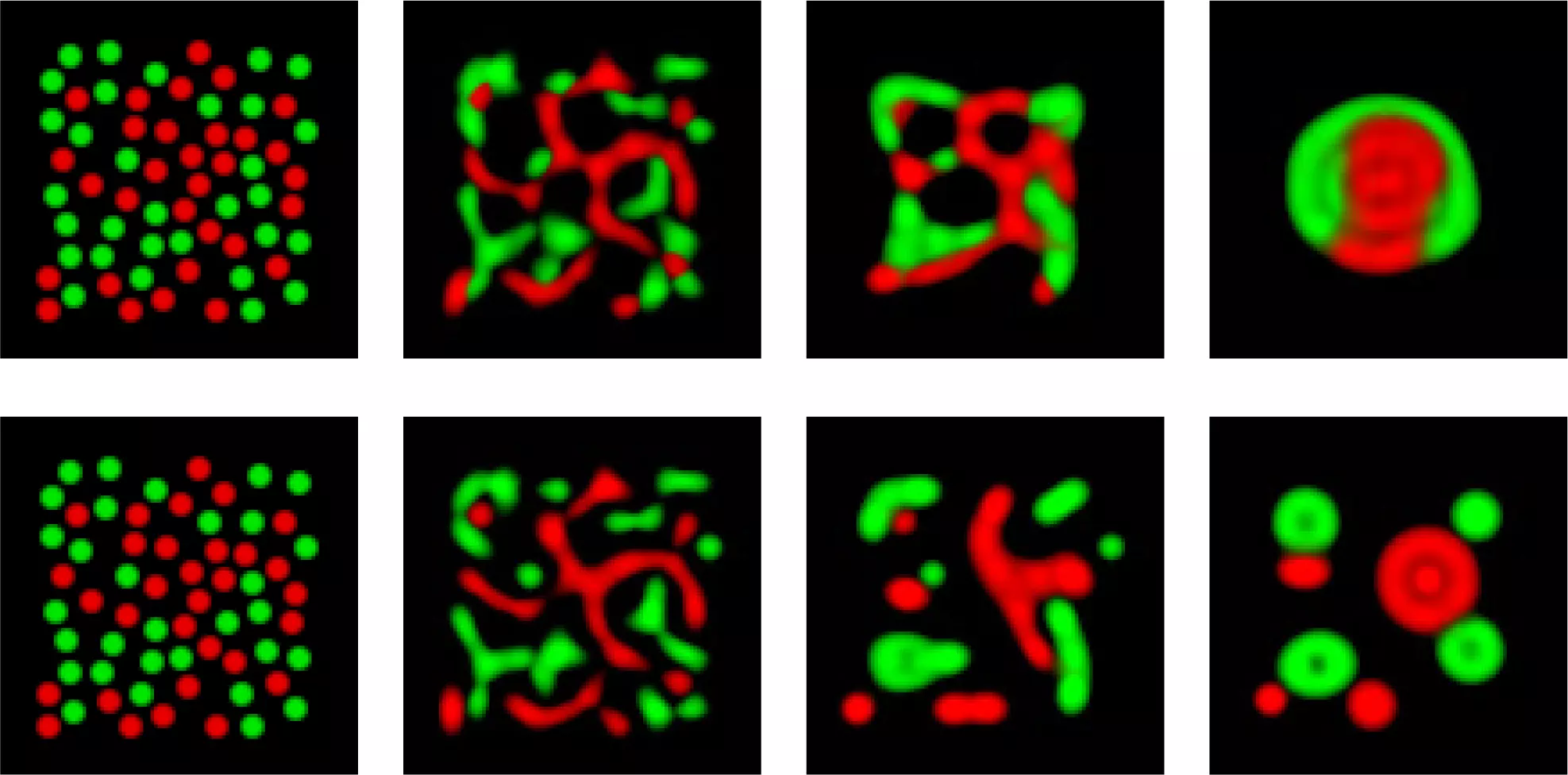

Computational Methods
We develop computational methods to simulate and infer dynamic models, discover novel elements, and find the best next experiments to test.
Ontologies and Databases
We create ontologies, curate databases, and develop expert systems used by both human scientists and artificial intelligence machines.
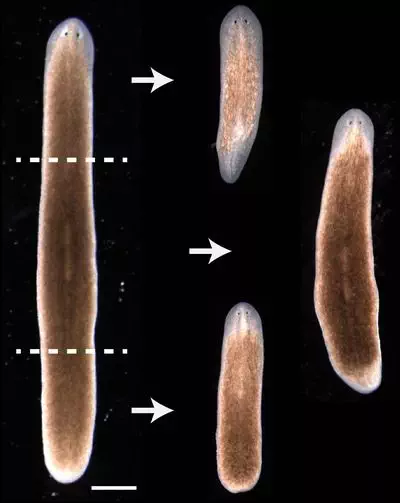
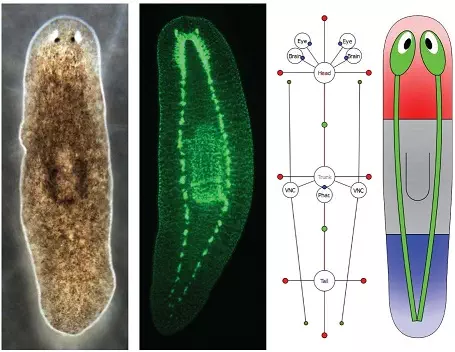
Development and Regeneration
We study how shapes and patterns are formed from a single cell during development and restored through regeneration.
Cancer and other Diseases
We seek to understand why and how regulatory mechanisms go awry to produce cancer and other diseases.
Synthetic Biology
We design and optimize regulatory and metabolic networks with desired dynamics and behaviors to solve specific bioengineering problems.



