Synthetic Biology
Automatic design of regulatory networks
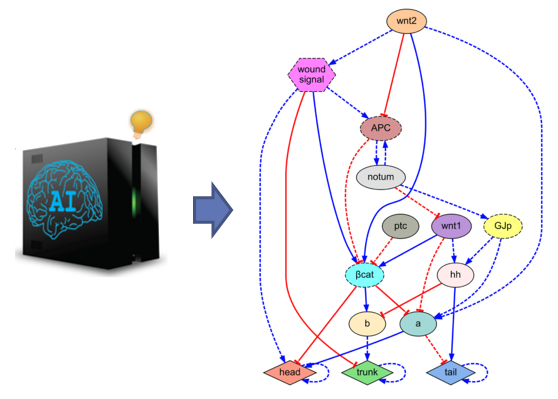
We are interested in software systems that automatically engineer regulatory networks that can perform and behave according to a desired function. The enormous possibilities of artificially-created genetic networks are currently limited to a few genes and simple functions due to the difficulty to design complex networks with many components and feedback loops. Using machines that can automatically design complex networks will pave the way for innovative engineering solutions to many current problems.
Publications
FROG analysis ensures the reproducibility of genome scale metabolic models
K. Raman, M. Kratochvil, et al., D. Lobo, et al., R.S. Malik-Sheriff
bioRxiv doi:10.1101/2024.09.24.614797, 2024.
Automatic design of gene regulatory mechanisms for spatial pattern formation
R. Mousavi, D. Lobo
NPJ Systems Biology and Applications 10, 35, 2024.
mergem: merging, comparing, and translating genome-scale metabolic models using universal identifiers
A. Hari, A. Zarrabi, D. Lobo
NAR Genomics and Bioinformatics 6(1), lqae010, 2024.
mergem: merging and comparing genome-scale metabolic models using universal identifiers
A. Hari, D. Lobo
bioRxiv doi:10.1101/2022.07.14.499633, 2022.
In situ probe and inhibitory RNA synthesis using streamlined gene cloning with Gibson assembly
A. Wolff, C. Wagner, J. Wolf, D. Lobo
STAR Protocols 3, 101458, 2022.
Inference of dynamic spatial GRN models with multi-GPU evolutionary computation
R. Mousavi, S.H. Konuru, D. Lobo
Briefings in Bioinformatics 22(5), bbab104, 2021.
Kinetic modeling of microbial growth, enzyme activity, and gene deletions: an integrated model of β-glucosidase function in Cellvibrio japonicus
J. Hwang, A. Hari, R. Cheng, J.G. Gardner, D. Lobo
Biotechnology and Bioengineering 117, pp. 3876-3890, 2020.
Fluxer: a web application to compute, analyze, and visualize genome-scale metabolic flux networks
A. Hari, D. Lobo
Nucleic Acids Research 48, pp. 427-435, 2020.
MoCha: molecular characterization of unknown pathways
D. Lobo, J. Hammelman, M. Levin
Journal of Computational Biology 23(4): 291-297, 2016.
Evolutionary development of tensegrity structures
D. Lobo, F.J. Vico
BioSystems 101(3), pp. 167-176, 2010.
Reconfiguration algorithms for robotically manipulatable structures
D. Lobo, D.A. Hjelle, H. Lipson
Proceedings of the ASME/IFToMM Intern. Conf. on Reconfigurable Mechanisms and Robots
J.S. Dai, M. Zoppi, X. Kong (eds.)
ReMAR2009 pp. 13-22, London, UK, 2009.