Planform can create, open, and modify databases stored in the edb file format. The following actions are available in the File menu:
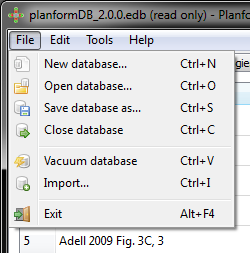
Planform main screen consists of several tabs that display the various types of data stored in a database. These include experiments, manipulations, morphologies, drugs, RNAi, species, and publications. In addition, the search tab allows querying the database for specific information. Clicking on a tab will bring up an alphabetic list of all current items corresponding to that data type in the database. Elements of each data type can be created, copied, or edited from these tabs directly, using the buttons on the bottom of the window. Double clicking an element will automatically bring up the edit screen, where all the details of the element can be viewed and edited.
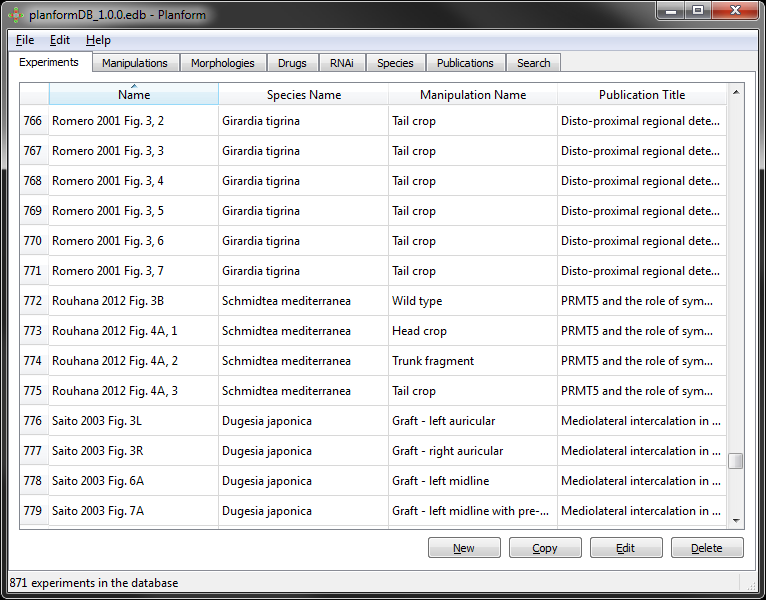
The search tab allows querying the database for searching specific data. A query is constructed with four parameters according to the following format:
The available operators for all the queries are the following:
Example: Search “Morphologies” with “Num. heads” “greater than” “1”. This search returns all morphologies that contain more than one head:
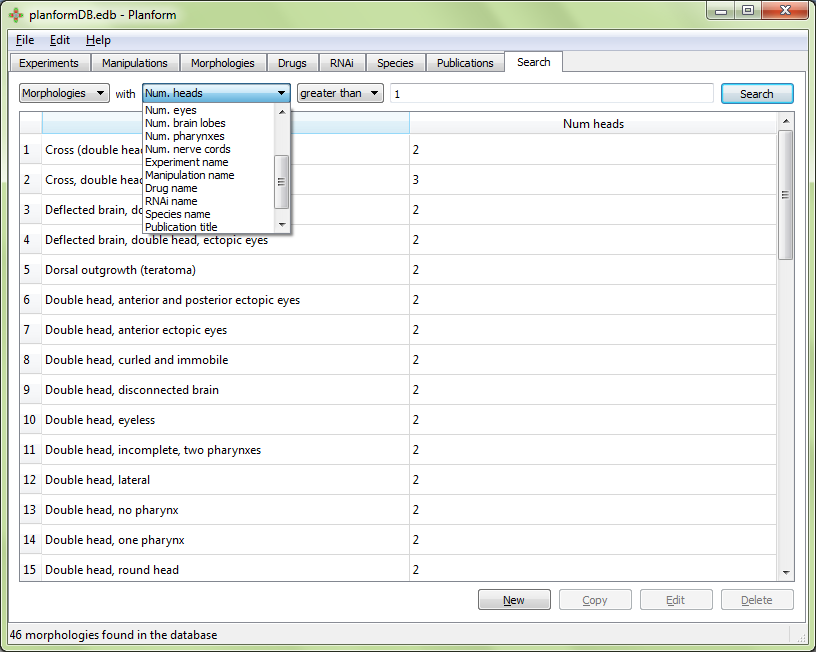
Each data type has specific characteristics that can be used in a query. These include:
Example: morphologies with RNAi equal to “Smed-bmp”. This would return all morphologies that can result from any experiment using Smed-bmp:

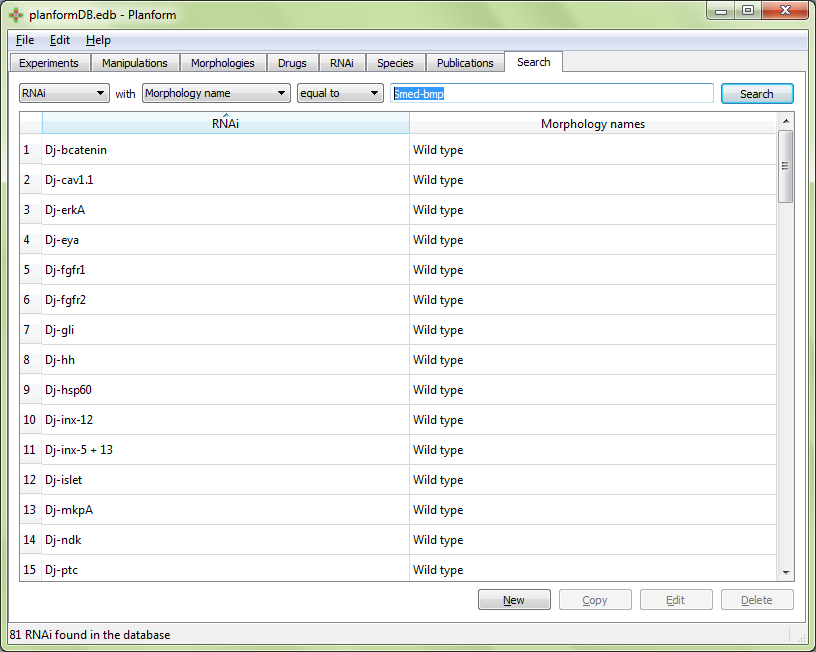
Example: RNAi with morphology name equal to “Wild Type”. This would return all RNAi that can result in the wild type morphology:
In addition, morphologies can be searched according to the number of the following features: regions, links, heads, trunks, tails, eyes, brain lobes, pharynxes, and nerve chords.
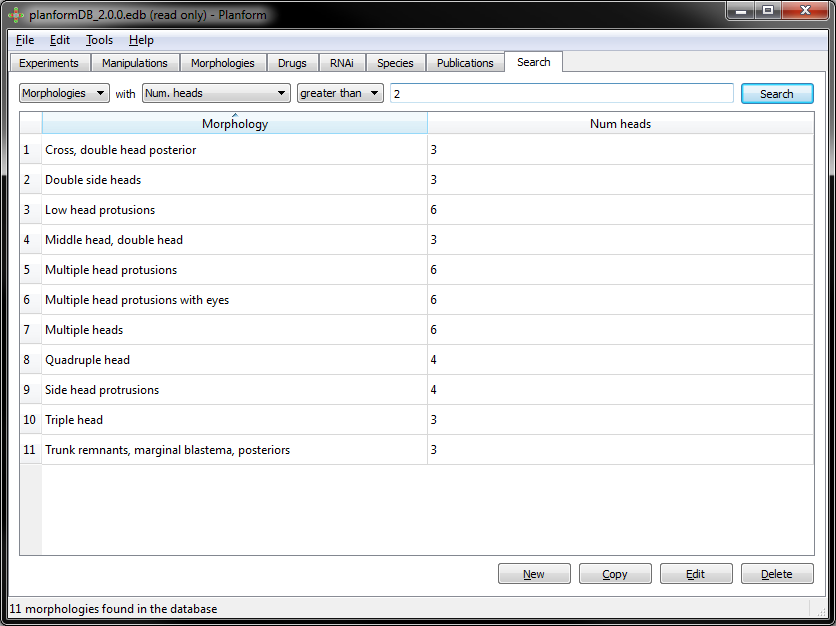
Example: Morphologies with number of heads greater than two:
Also, publications can be searched by either title or year.
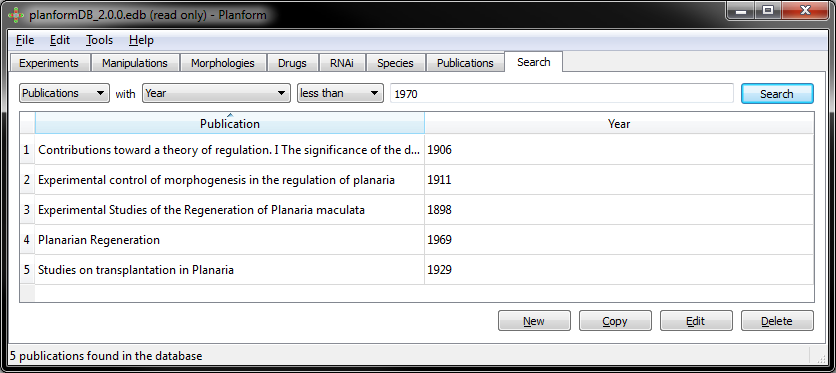
Example: Publications from earlier than 1970:
Finally, a quick search can be performed from any tabs (including the search results). Right clicking on an element will give the option to search for any other parameter with that element as the limiting parameter.
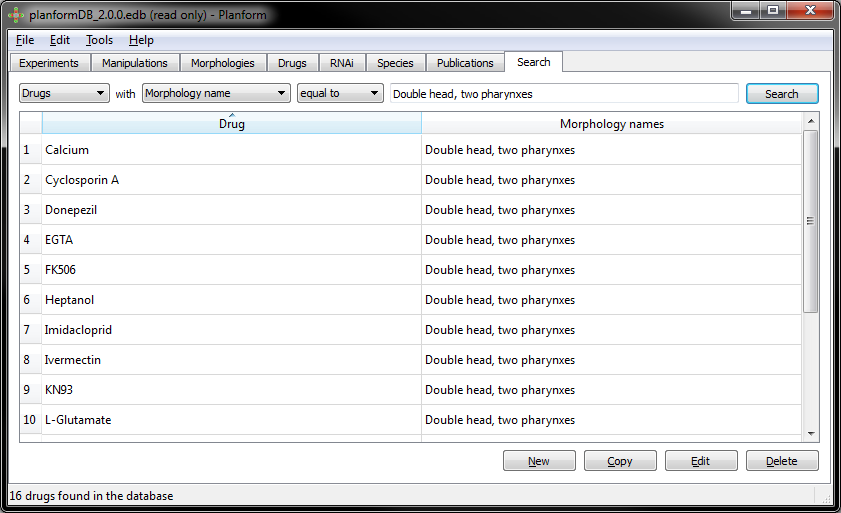
Example: Searching all the drugs that produce a “double head, two pharynxes” morphology:
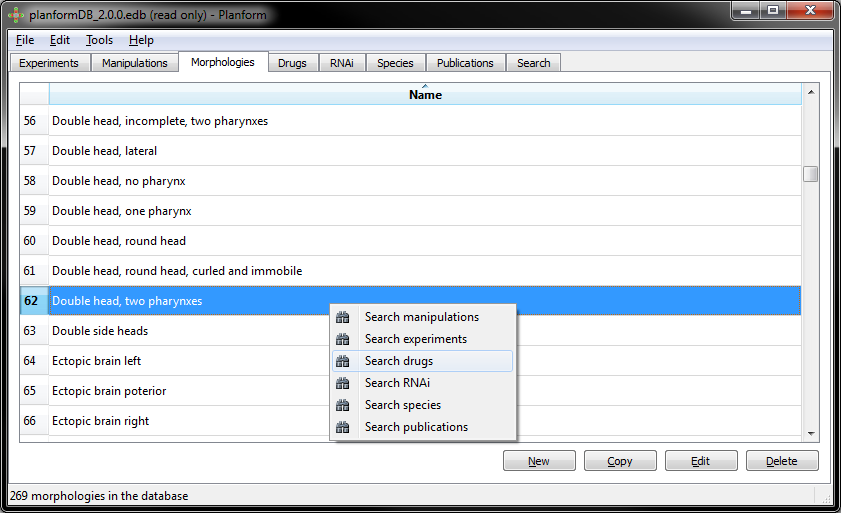
16 different drugs produce a “double head, two pharynxes” morphology:
A new experiment can be created from scratch or copied from an existing experiment by clicking on “New” or “Copy” at the bottom of the experiment tab in the main window. Double-clicking on an experiment or selecting “Edit” in the main window edits the experiment. During the creation or edition of an experiment, the following interface shows the characteristics of the experiment:
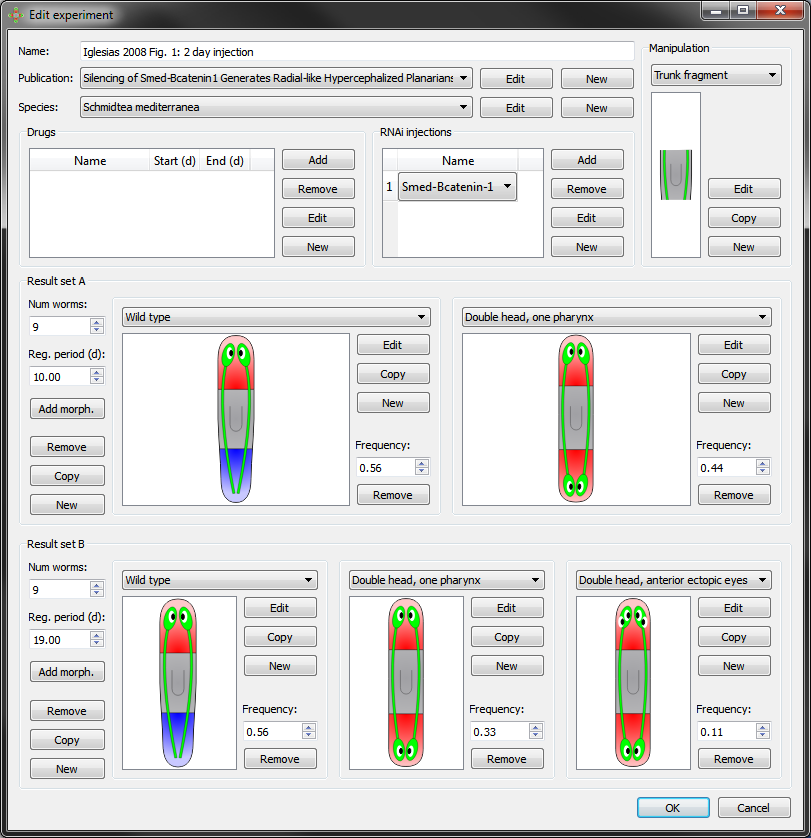
An experiment is divided into its general information, procedure, and results.
The first information to be input in an experiment are the name of the experiment (see naming conventions), the publication, and the species of worm used in the experiment. The publication and species can be selected from an existing one using the dropdown menu or a new one can be created by clicking on “New”. An existing publication or species can also be edited from this screen.
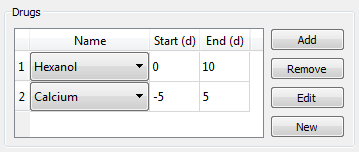
The experiment procedure consist of three parts: drugs applied, RNAi injected, and the worm manipulation.
Drugs are defined in two ways: the drug used and the time period during which it was used. Drugs can be added, removed, edited, or created from scratch by clicking on the available buttons in the interface. Once a drug is added to the experiment, a dropdown menu will allow the selection of the specific drug. The starting and ending times define when the drug was used, being the time relative to the manipulation of the worm (time 0). Zero is the default starting and ending time if no times are given. A drug with negative time means that it was administered before performing the manipulation. When relevant, dosage information can be included in the title of the experiment.
Many different RNAi are typically used in planarian regenerative experiments. Similar to drugs, RNAi can be added removed, edited, or created from the experiment interface. For RNAi, no time information is directly included, since it is not usually relevant for the results. Any timing or dosage information relevant for the experiment can be included in the title of the experiment.
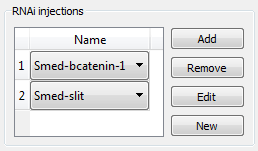
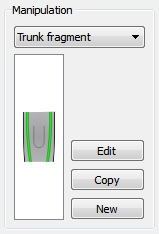
A specific worm manipulation can be specified for an experiment. If no manipulation has been made, the wild type morphology can be selected as “no manipulation”. A previously defined manipulation can be selected from the dropdown menu, or a new one can be created. Clicking in the corresponding button, the manipulation editor can be opened from this screen in order to edit, copy, or create a new manipulation for the experiment (see manipulations).
The results of an experiment are specified with result sets. Each result set defines the outcome (resultant morphologies) for a given number of worms after a specific regeneration period since the manipulation. Result sets can be removed, copied, or created from scratch to describe the result of an experiment over different time periods or in different trials. If a different manipulation is used, a separate experiment must be made.
Each result set contains several morphologies that correspond to the phenotypes observed in the experiment. Morphologies can be added or removed from the result set. A dropdown menu allows the selection of each phenotype, which can then be edited or copied, or a new morphology can be created from scratch (see morphologies). Finally, a frequency out of 1 for each morphology defines the penetrance of each of the morphologies in the specific result set. The sum of frequencies must add up to 1. If no frequencies are given, equal amounts are assumed.
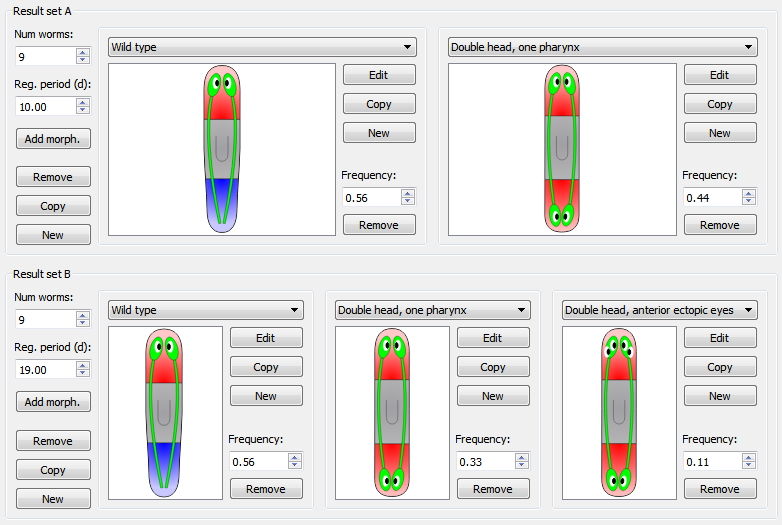
The following example shows the results from an experiment. Two different result sets were obtained. The first (result set A) specifies the resultant morphologies of 9 worms after 10 days: 56% (5 worms) regenerated the wild type morphology, whereas 44% (4 worms) regenerated a double head, one pharynx phenotype. The second (result set B) specifies the resultant morphologies of another (or same than before) 9 worms after 19 days: 56% (5 worms) regenerated the wild type morphology, 33% (3 worms) regenerated a double head, one pharynx morphology, and 11% (1 worm) regenerated a morphology characterized by a double head and anterior ectopic eyes:
Manipulations are created and edited in the manipulation editor, which is divided into two panels. The left panel shows the graphical formalization (defined with a tree structure) of the manipulations being performed. The right panel shows a cartoon representation to aid in the visualization of the manipulation.
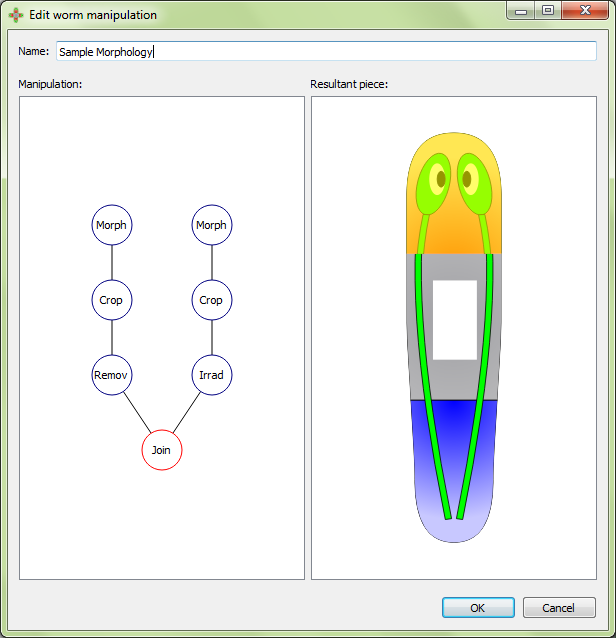
To add a manipulation action, left click on the previous manipulation action and then select the type of the new manipulation to add. A manipulation can be added between two existing actions by clicking the first one. Finally, a manipulation can be also deleted by selecting the appropriate action after right clicking on it:
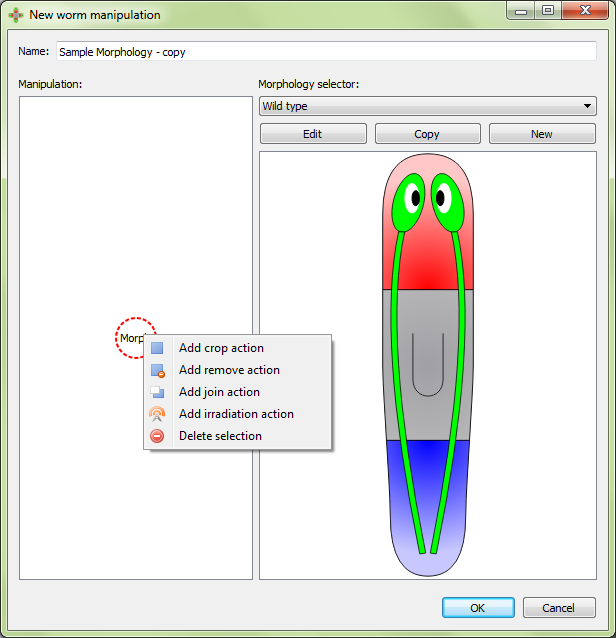
The type of manipulation actions are described below.
When starting a new manipulation, the first selection to be made is the morphology of the starting worm. Any existing morphology can be used or a new one may be created. To change the morphology, click on the morphology marker on the left half of the panel, and then select the desired morphology from the drop down menu on the right:
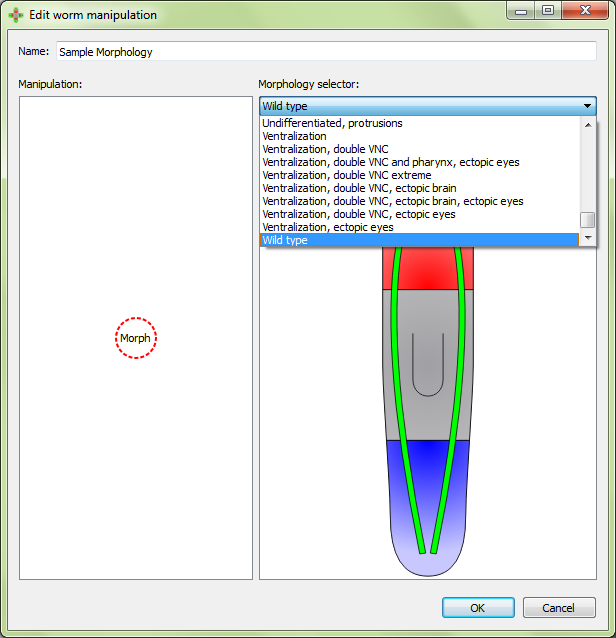
Three types of manipulations are defined by a single polygonal area: crop, remove, and irradiation. Selecting one of these manipulations actions will produce a box in the right pane. Drag the vertices of the box so that it covers the desired area. Left clicking then right clicking on a vertex will give the option to delete or add a vertex.
Crop will keep only the area inside of the polygon:
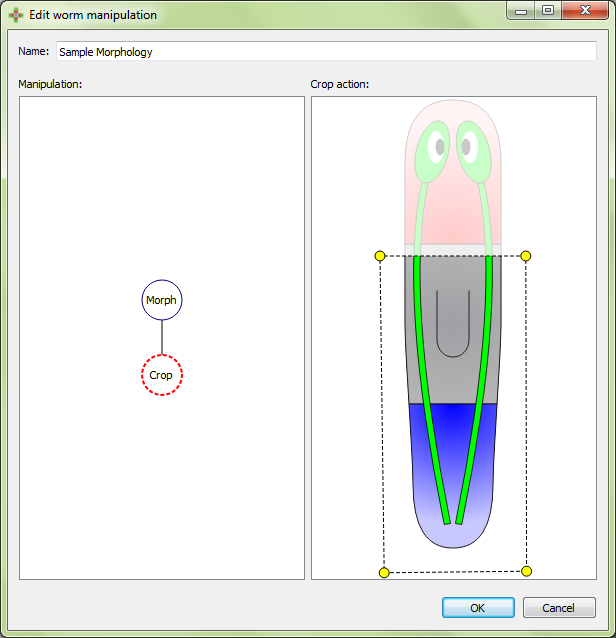
Remove will keep only area outside of the polygon:
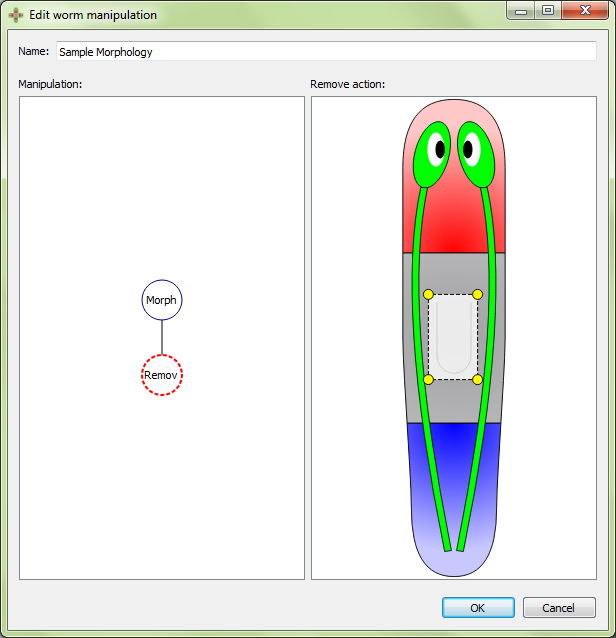
Irradiate represents irradiation within the polygon (the doses of the radiation are usually recorded in the manipulation name):
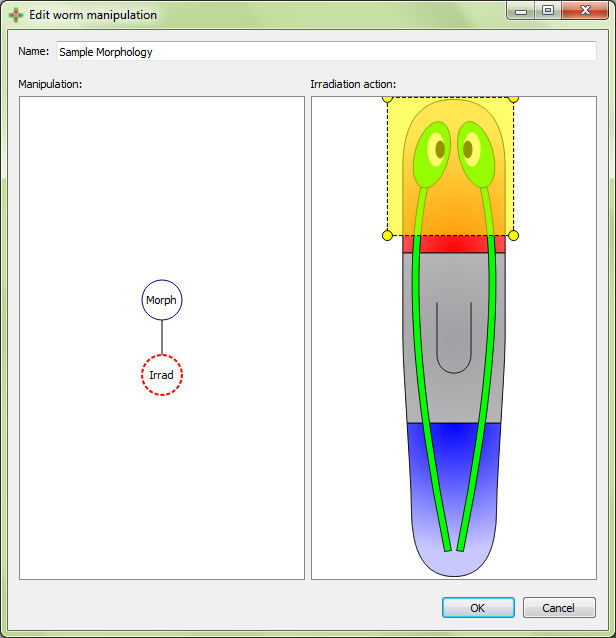
The join manipulation action corresponds to grafting two worms or parts of worms together. Selecting a join action will produce a second branch in the manipulation tree. Here, a second morphology can be selected, along with any additional manipulation actions desired. The left tree represents the fixed piece; the right tree represents the movable piece. Once all desired manipulations have been added to the join tree, click on the join action marker in the left panel and both pieces will then appear in the right panel. Clicking and dragging inside the movable piece in the right panel will control movement of the piece. Clicking and dragging outside the movable piece will control the rotation of the piece.
The join manipulation bifurcates the manipulation tree:
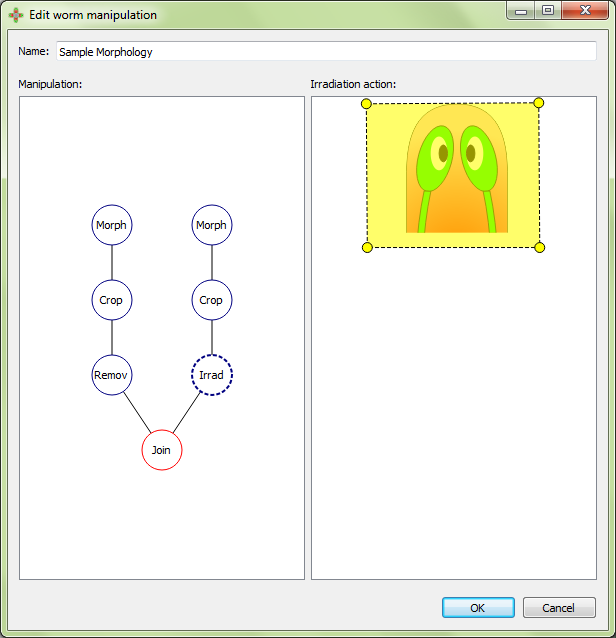
The graft position is adjusted in the interface:
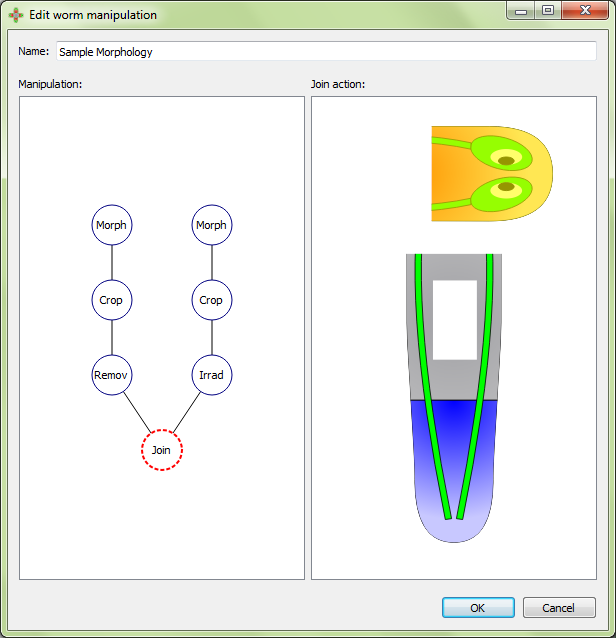
The graft position is adjusted in the interface:
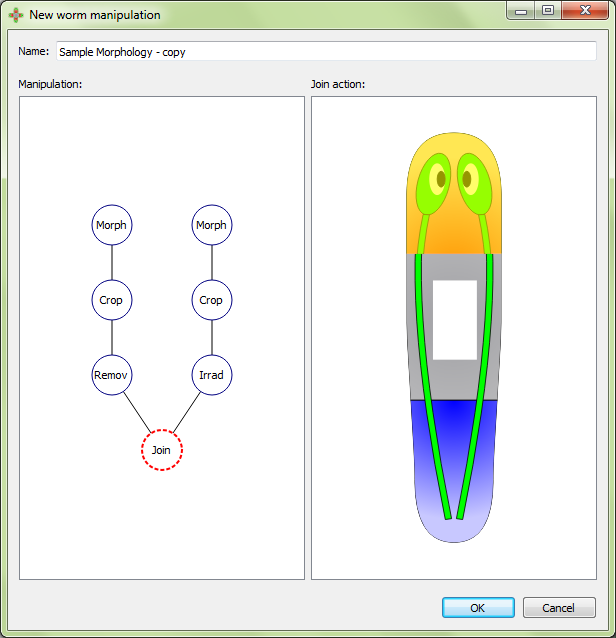
Morphologies are created and edited in the morphology editor, which is divided into two panels. The left panel shows the graphical formalization of the morphology, while the right panel shows a cartoon representation to help with the visualization of the morphology being created.
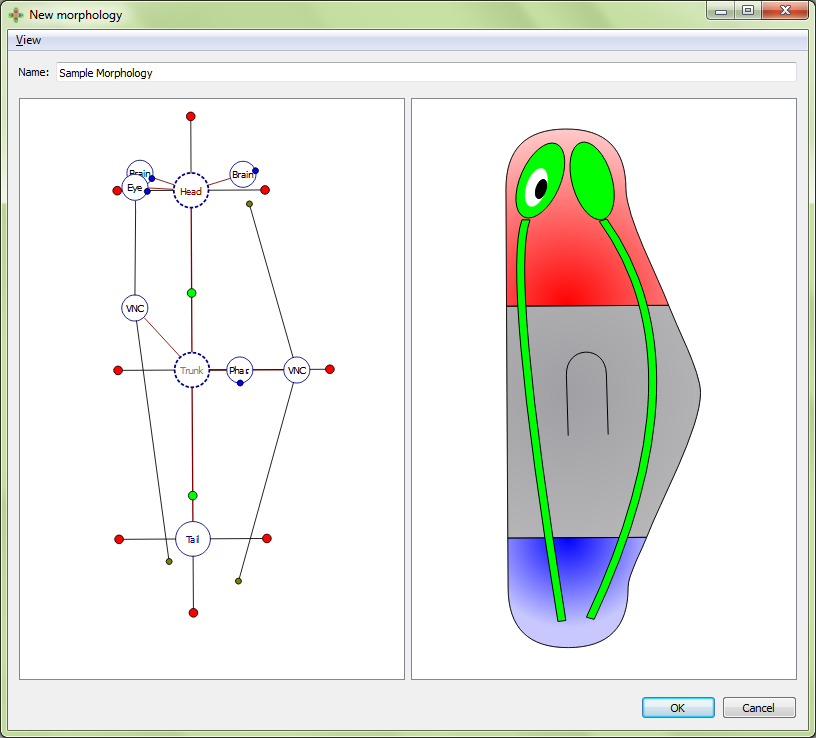
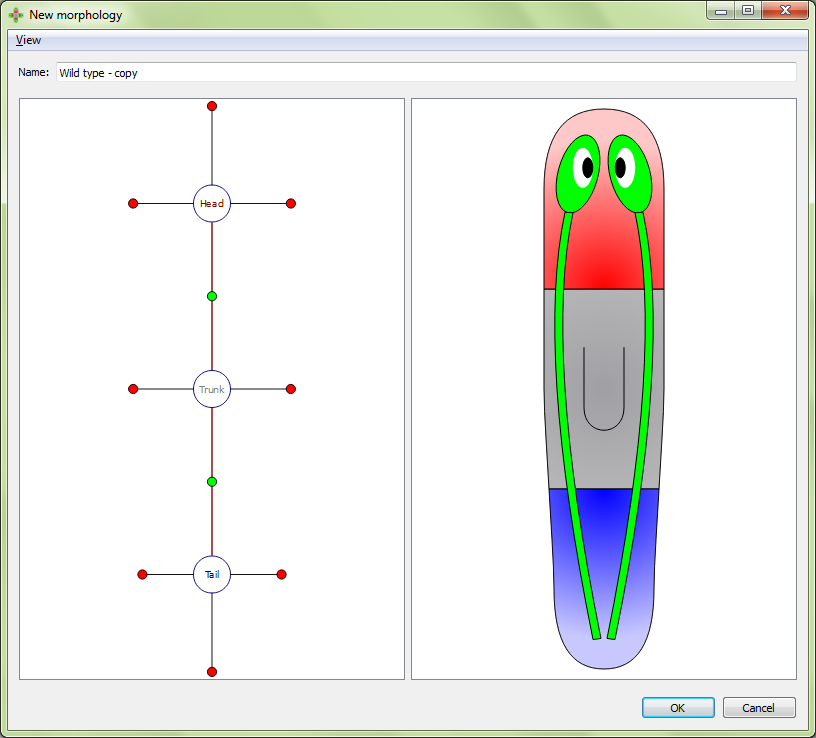
New morphologies are usually created by copying and editing the wild type morphology. However, new morphologies can be created from scratch also, as we describe next.
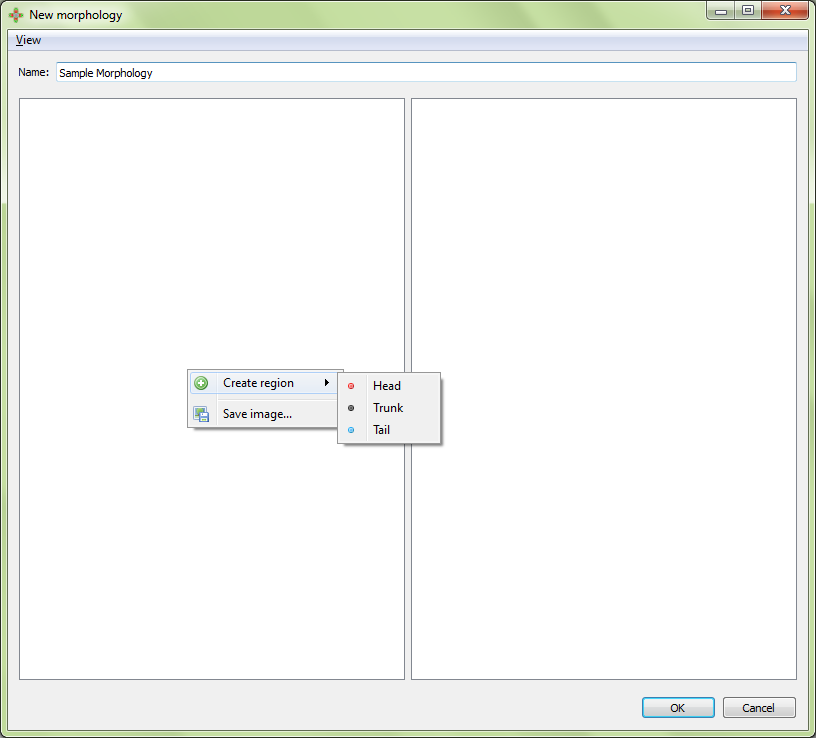
The first step of creating a morphology is to define the regions it contains. These include head, trunk, and tail regions. To add a first region, right click on the left panel and select the desired region under “Create region”:
Left clicking on an existing region will provide several options:
Add region: adds a new region with a connection to the selected region.
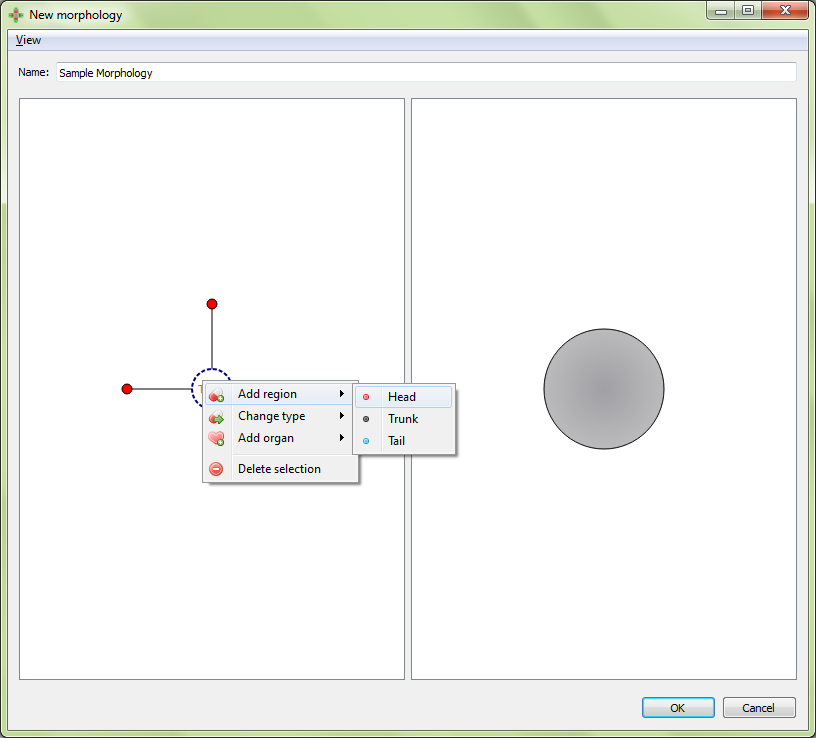
Change type: changes the region type of the selected region.
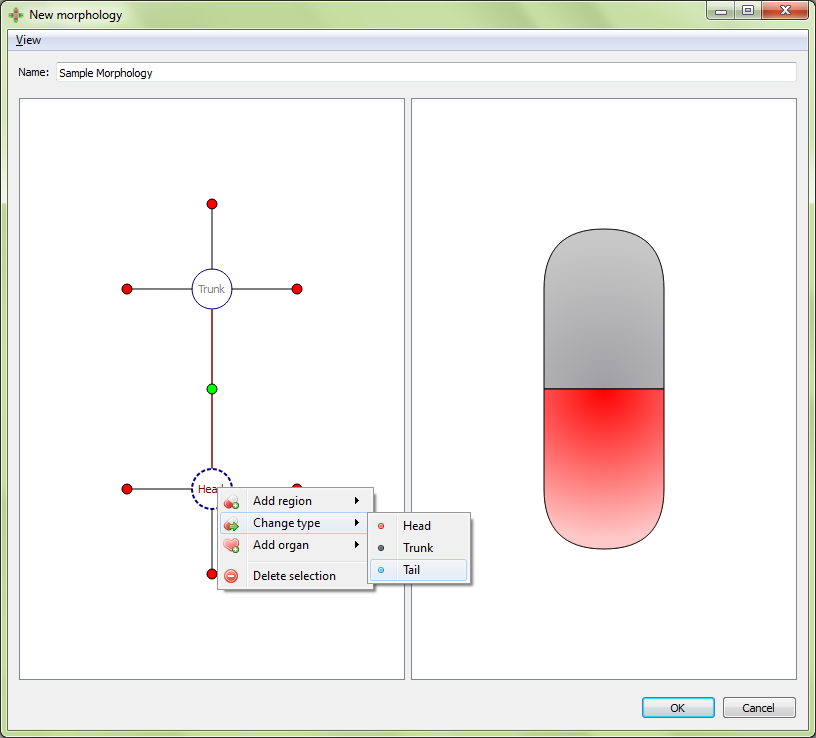
Result:
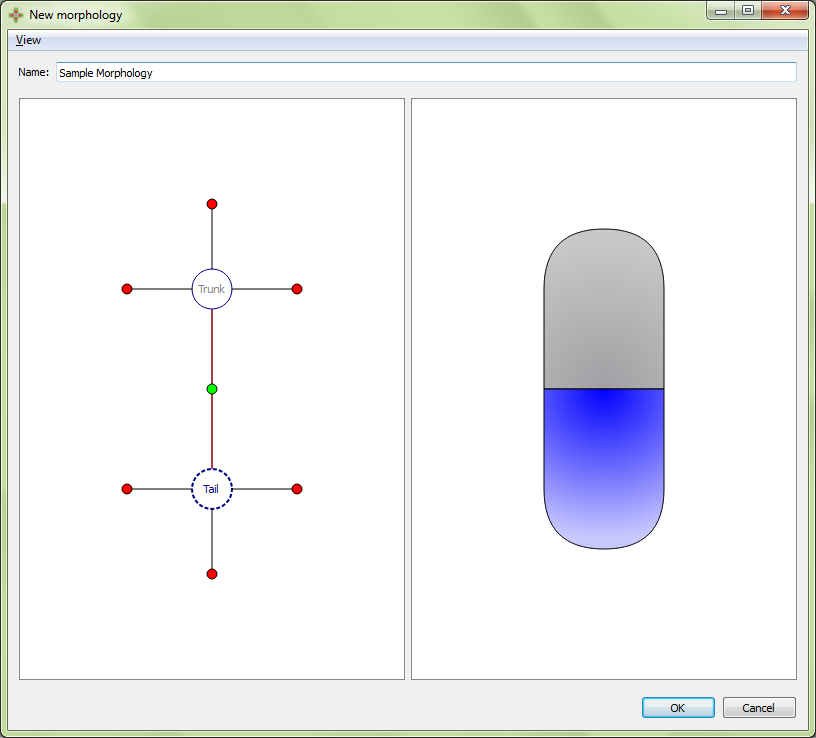
Add organ: adds a new organ to the selected region. See below for details.
Delete selection: deletes the selected region. This also deletes any organs in that region.
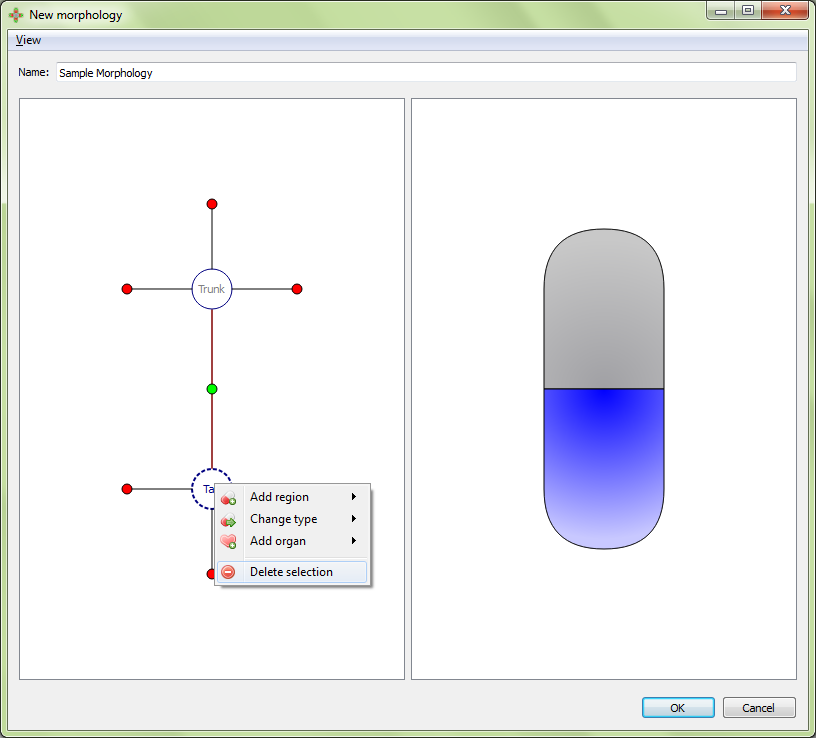
Result:
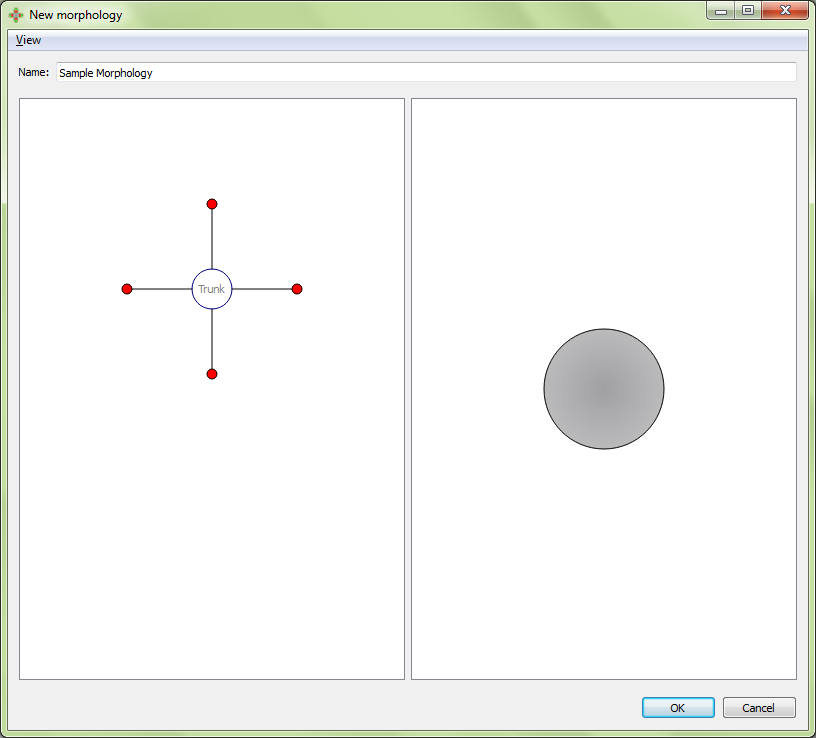
Any number of any types of regions may be added to the morphology in any combination. Regions can be moved to the desired place by clicking and dragging the marker in the left panel.
Every region is defined by a number of external boundaries and connection boundaries. A region connected to a single other region will have three external boundaries. A region connected to two or more other regions will have an external boundary between every set of connections. There is one connection boundary defining every pair of connected regions. To adjust either type of boundary, left click and drag the colored circle corresponding to the boundary. External boundaries are defined by red circles, while connection boundaries are defined by green circles. External boundaries define the shape of the region itself. Connection boundaries define where one region stops and the next begins.
Initial morphology:
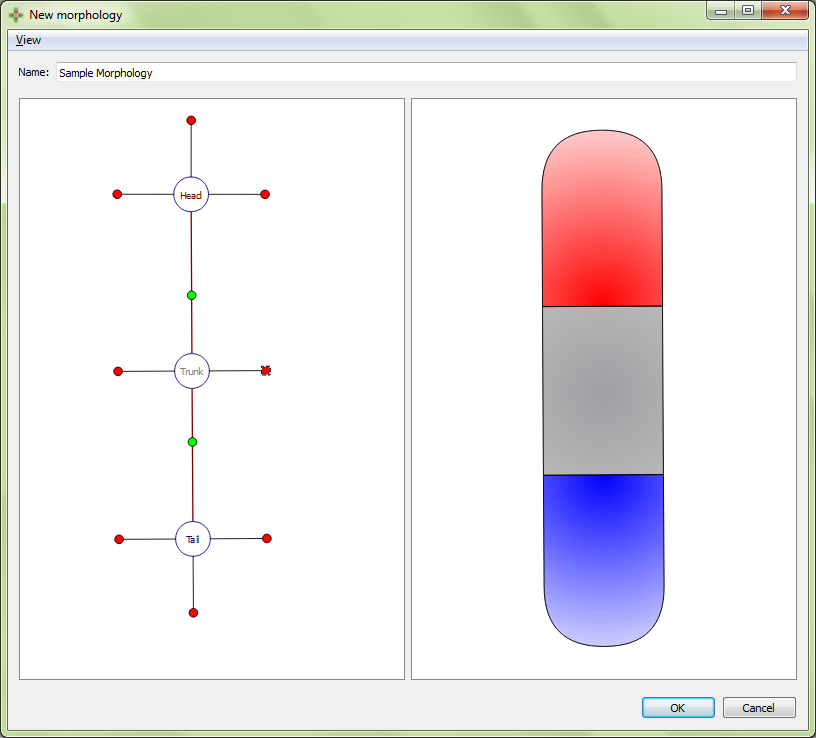
Adjusting region-region boundary:
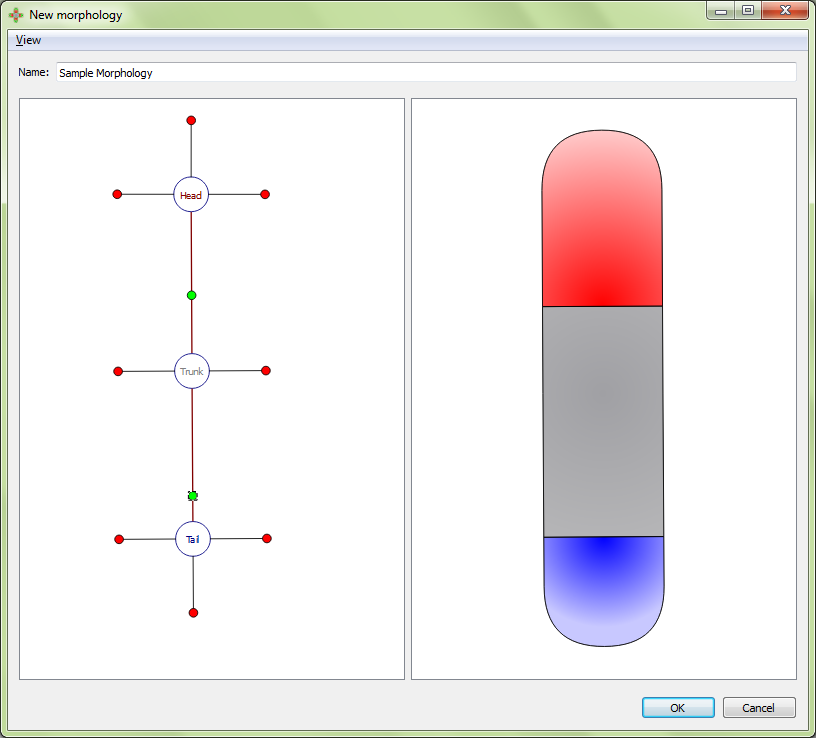
Adjusting external boundary:
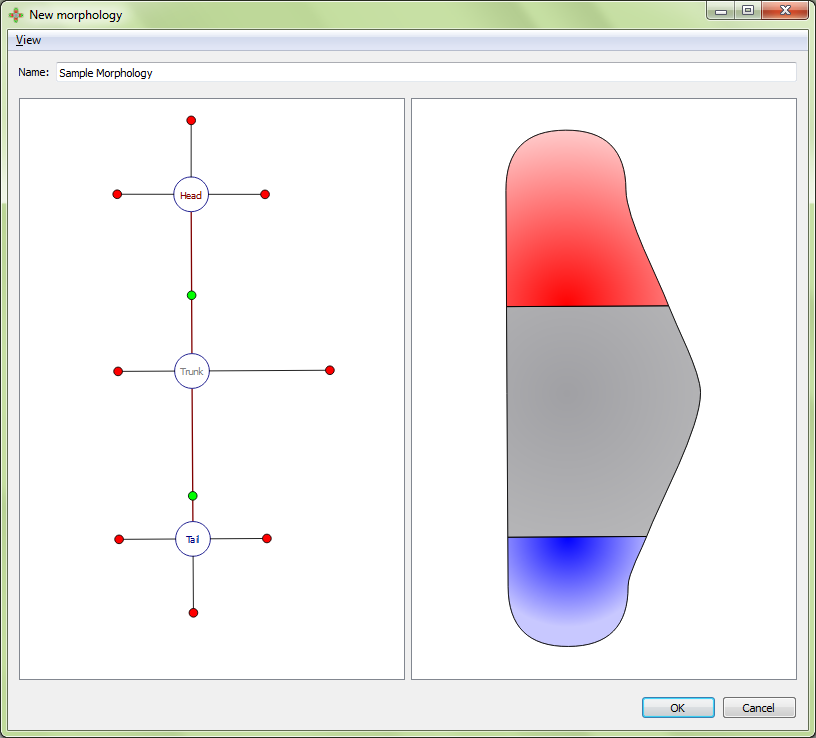
Organs can be added to the selected region by right clicking the region marker in the left panel and selecting the desired organ type under “Add organ”. Organs include eyes, brains, pharynxes, and ventral never cords (VNC). Any organ may be added to any type of region. The wild type worm has two eyes and two brains in the head, a pharynx in the trunk, and two VNC’s in the trunk that span the length of the worm. To delete the selected organ, right click on the organ’s marker in the left panel and choose “delete selection”.
Adding an organ to a region:
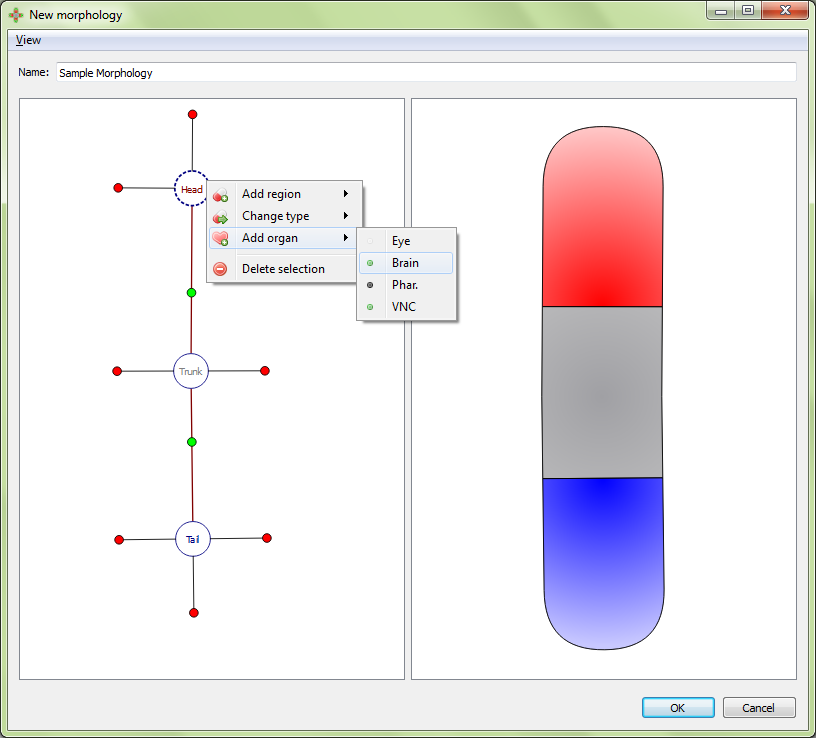
Resultant morphology:
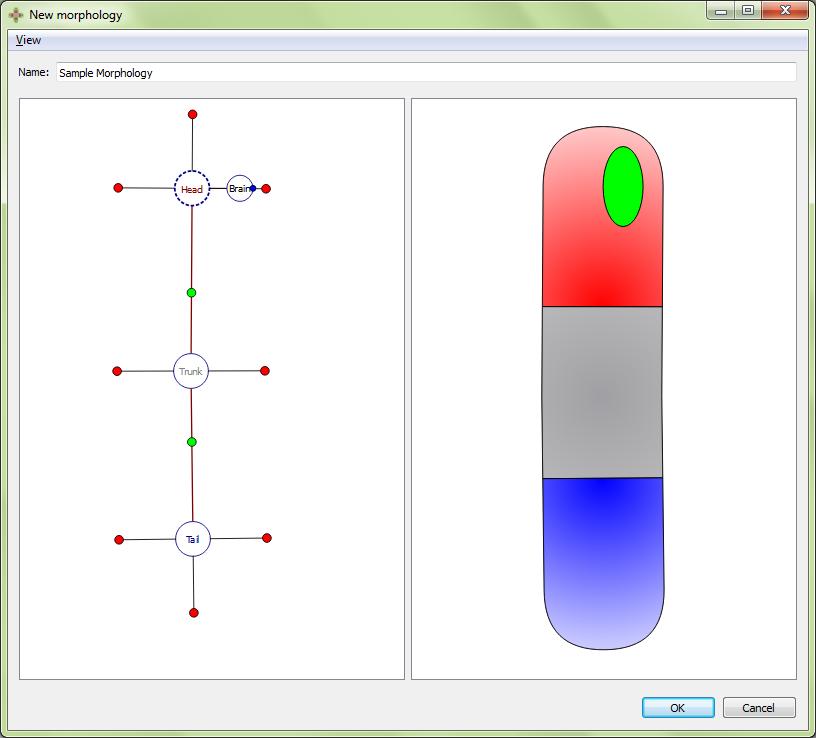
Organ markers will only appear when its region has been selected first; however, organ markers can be set always visible by activating the “All parameters” option in the View top menu.
Morphology view with all parameters option off:
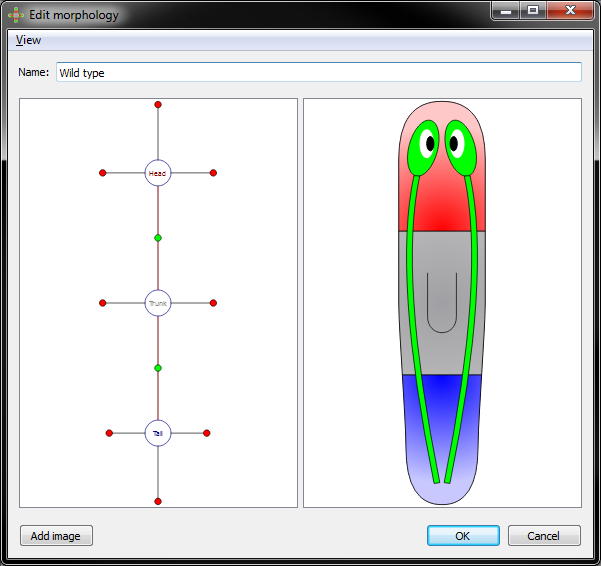
Morphology view with all parameters option on:
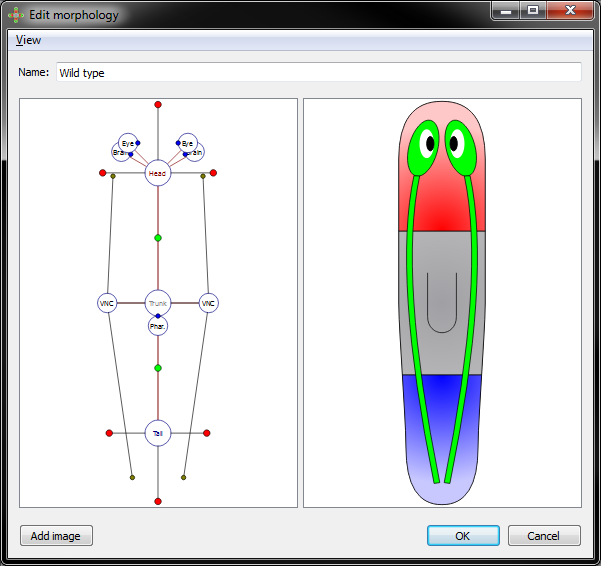
Eye, brain, and pharynx organs are defined by a location and a rotation. To move the location of an organ, left click and drag its marker to the desired location. To rotate the selected organ, left click and drag the blue circle on the border of the marker.
Displacing and rotating an organ:
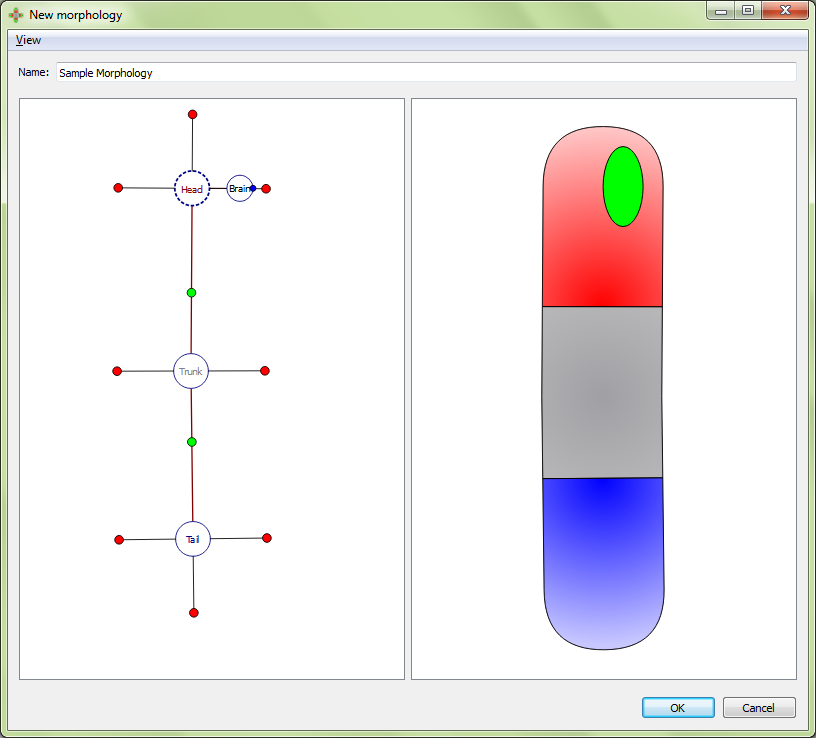
Resultant morphology:
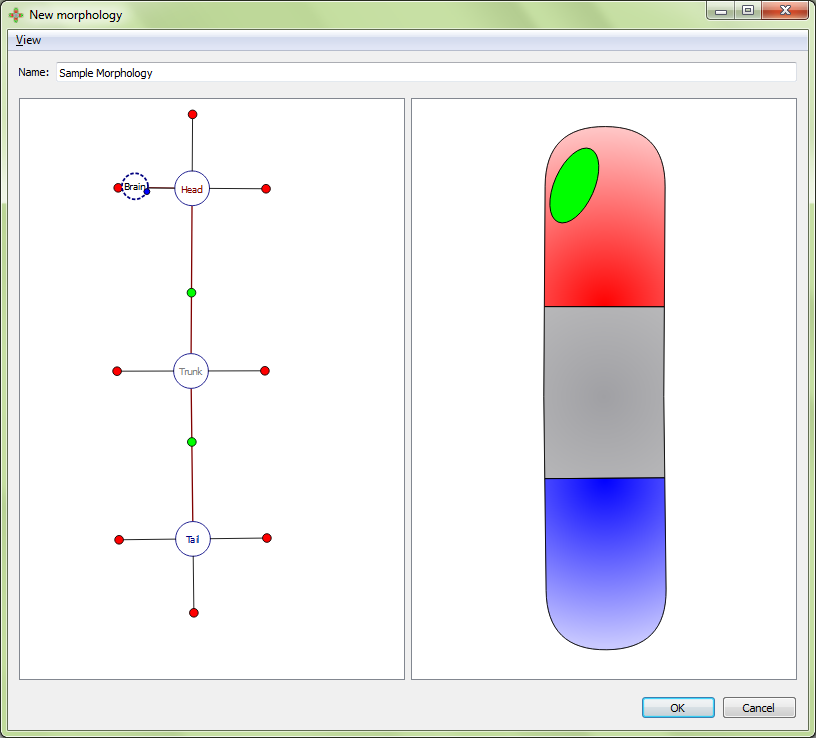
A VNC is defined by a curve made of three points. To adjust the location of the curve points, select the organ’s marker, and then click and drag either the marker (the middle point defining the curve) or one of the two olive-color circles connected to the marker (the end points defining the curve).
Adding a VNC:
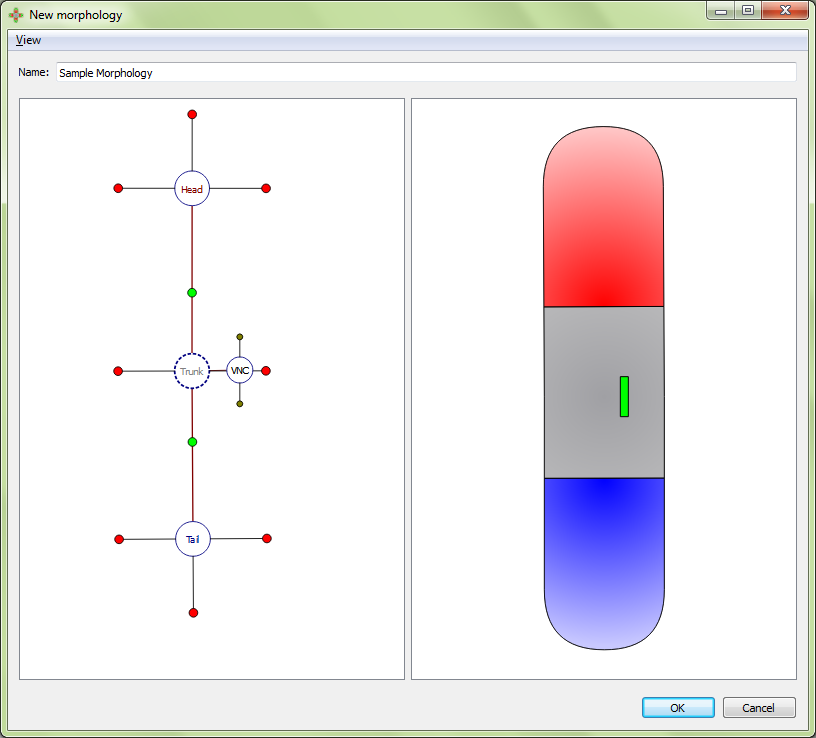
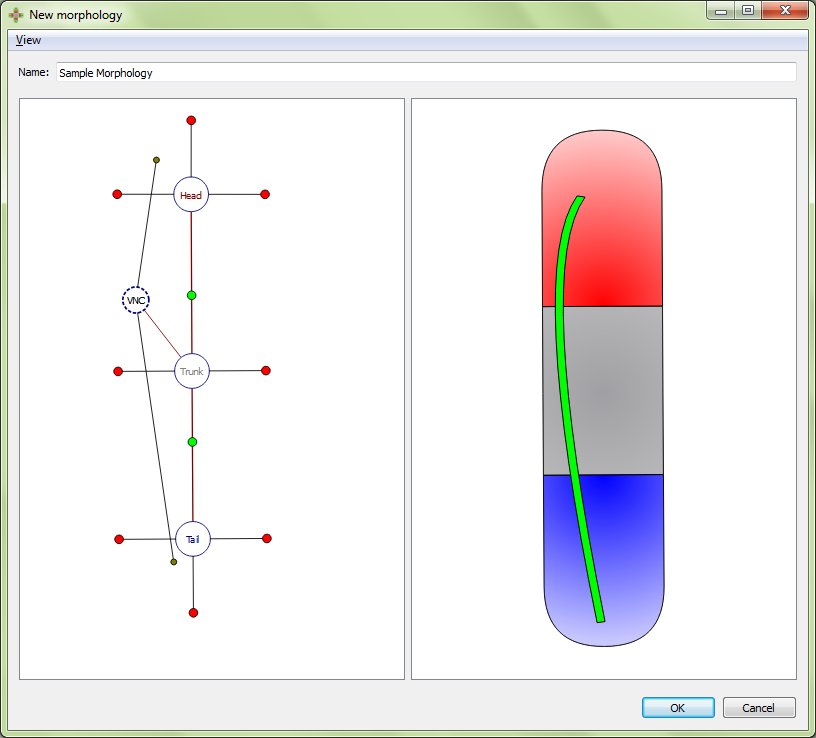
Adjusting a VNC:
Any names may be used in the database, but several conventions were used when making the central database.
The following rules were used for naming experiments in the database:
Manipulations and Morphologies names are created as explicit and modular as possible. Manipulation names describe each type of manipulation action performed in succession. Morphology names describe sequentially each morphological variation from the wild type. The goal of using a modular approach is to aid in both locating existing entries and adding new entries in an organized fashion. Common traits are always put first in names, so that similar manipulations and morphologies are in close proximity in alphabetical order.
RNAi are named based on the species and the encoded protein knocked out.
Clicking off the image in either panel in an editor, and then holding ctrl and sliding the mouse wheel will zoom the contents of the screen in or out. Alternatively, when using a laptop pad, zooming can be achieved by holding one finger fixed and sliding a second finger.
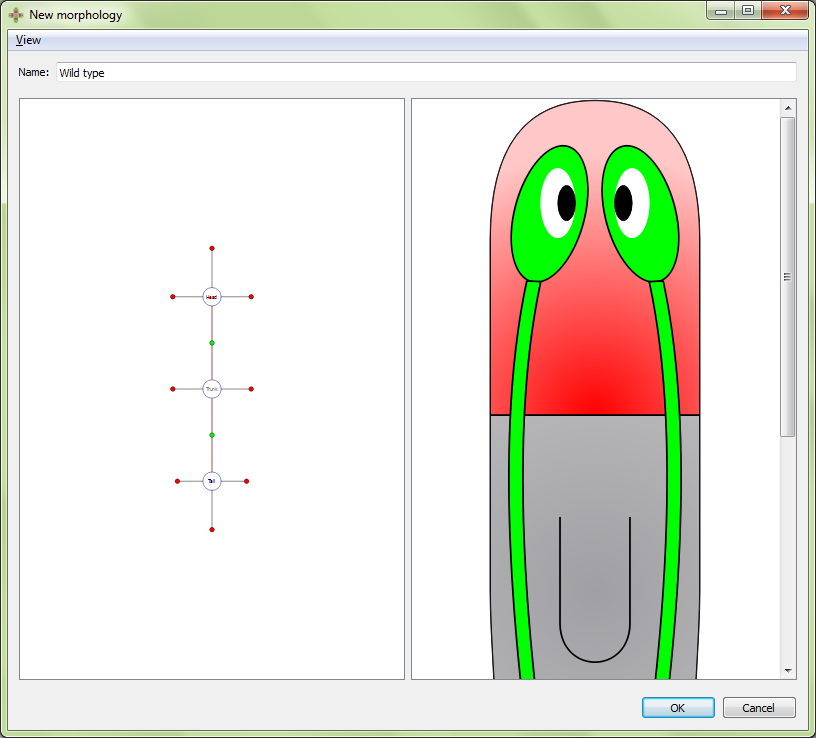
Right clicking on the blank area of any image will give the option to export the image as a png, jpg, or bmp file: