MoCha
Molecular Characterization of Unknown Pathways
Summary
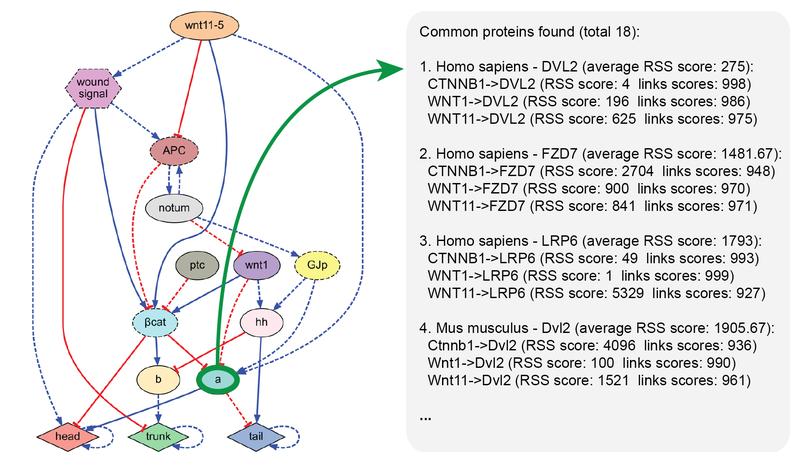
MoCha (Molecular Characterization) is an efficient tool for the characterization of unknown components and pathways interacting with a a given set of proteins. MoCha can search within seconds over one billion protein-protein interactions curated in the STRING database. MoCha and its source code is freely distributed in this website.
Update April, 2020
MoCha has been updated to work with the latest version of the STRING database (v11). Now MoCha searches for pathway interactions in more than 5,000 organisms, 14 million proteins, and 6 billion links. To efficiently process this very large dataset within seconds, MoCha needs of 430 Gigabytes of disk space for its installation.
Software
You can freely use and download MoCha and its source code (GPLv3 license) in your own computer. A script (Unix/Linux) is included to automatically download the necessary database files, process them, and compile the program for your system. Please, click here to download the latest version of MoCha.
Citation
MoCha: molecular characterization of unknown pathways
D. Lobo, J. Hammelman, M. Levin
Journal of Computational Biology 23(4): 291-297, 2016.
Acknowledgments
This work was supported by National Science Foundation (EF-1124651), National Institutes of Health (GM078484), W. M. Keck Foundation, and G. Harold and Leila Y. Mathers Charitable Foundation. Computation used equipment awarded by Silicon Mechanics.